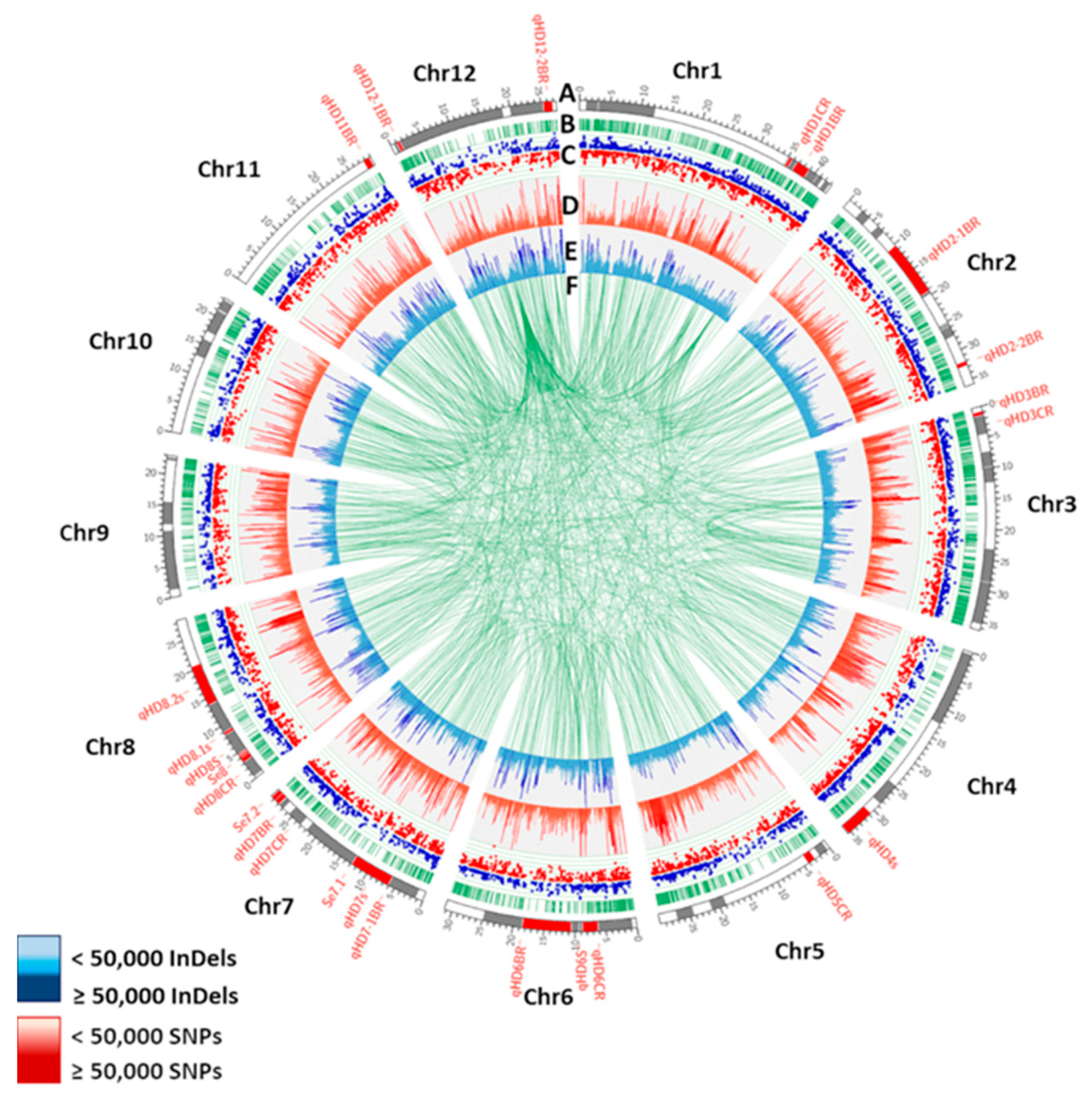
IJMS | Free Full-Text | Whole-Genome Sequencing and RNA-Seq Reveal Differences in Genetic Mechanism for Flowering Response between Weedy Rice and Cultivated Rice
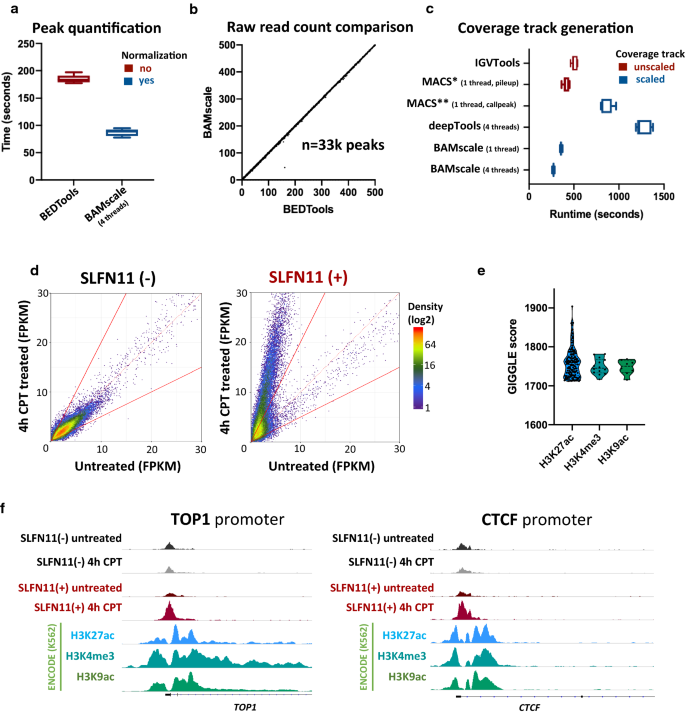
BAMscale: quantification of next-generation sequencing peaks and generation of scaled coverage tracks | Epigenetics & Chromatin | Full Text

SkewC: Identifying cells with skewed gene body coverage in single-cell RNA sequencing data - ScienceDirect
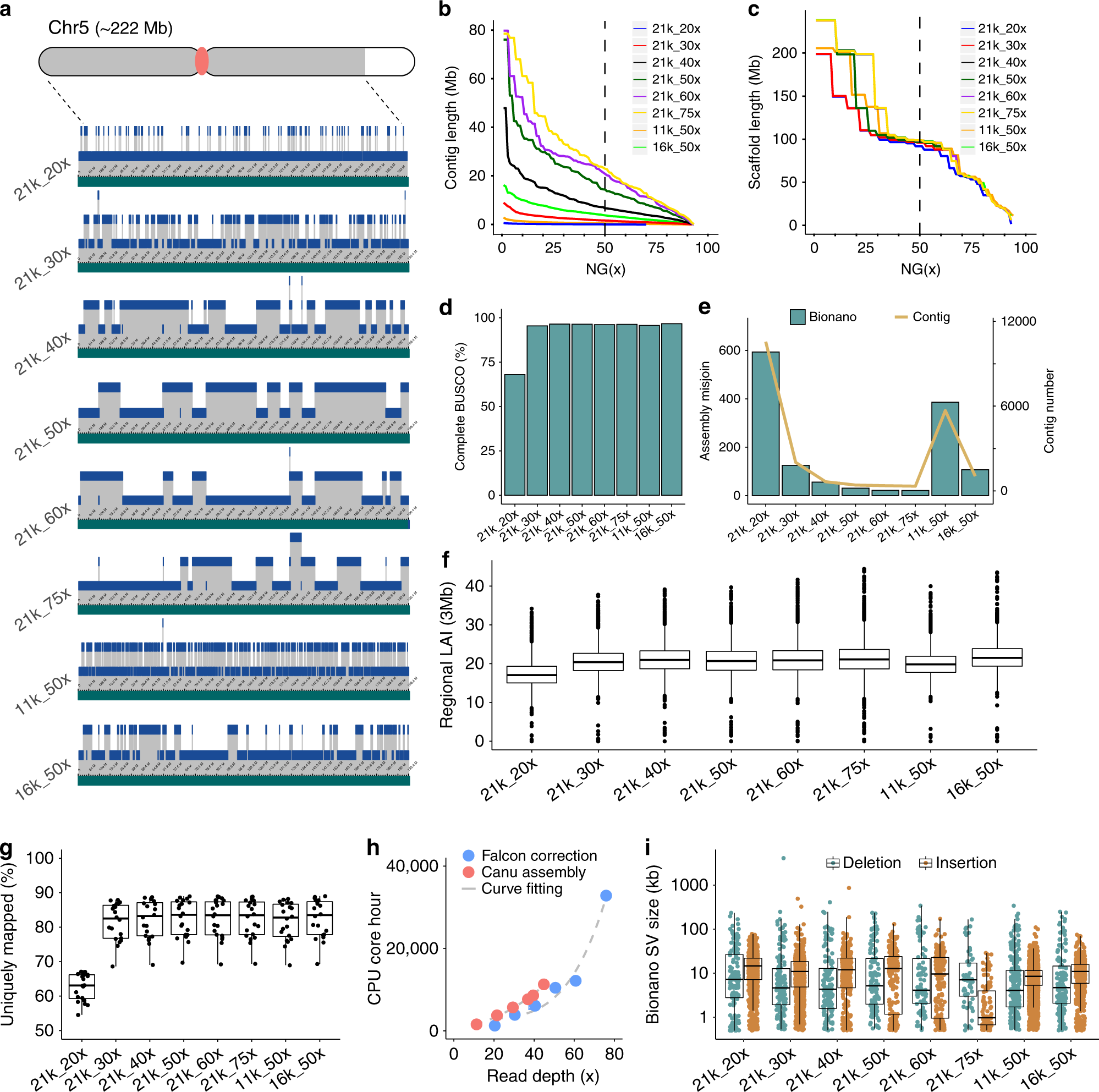
Effect of sequence depth and length in long-read assembly of the maize inbred NC358 | Nature Communications
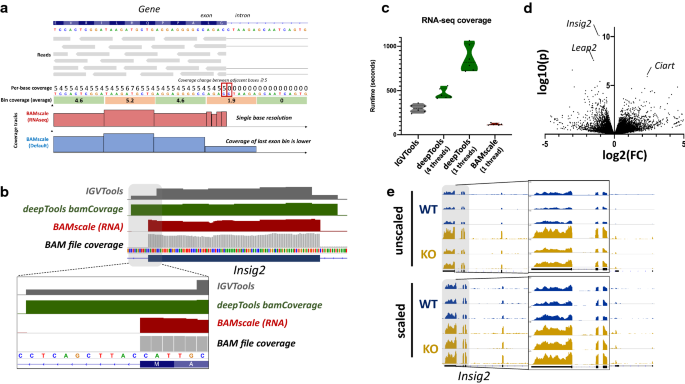
BAMscale: quantification of next-generation sequencing peaks and generation of scaled coverage tracks | Epigenetics & Chromatin | Full Text
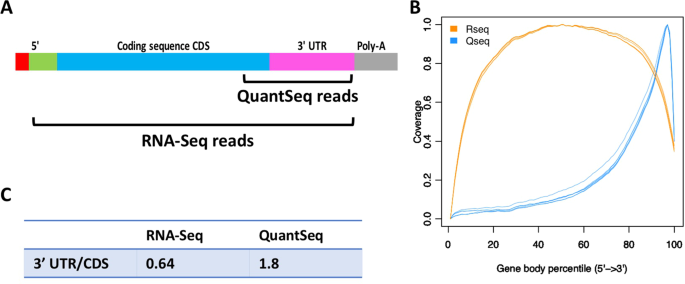
QuantSeq. 3′ Sequencing combined with Salmon provides a fast, reliable approach for high throughput RNA expression analysis | Scientific Reports